install.packages("rstatix")5 R packages
When we have finished this chapter, we should be able to:
5.1 What are R packages?
5.1.1 Standard (base) R packages
R installs a set of standard (base) packages during the installation process. Standard packages are stored in a library folder of the R program and contain the basic functions that allow R to work as well as a number of statistical and graphical functions that are ready to be used in our data analysis.
5.1.2 Add-on packages
More packages can be added later in User’s R library from repositories, when they are needed for some specific purpose (add-on R packages). Add-on R packages are created by a world-wide active community of developers and R users, covering a wide number of different applications. Most of these packages can be installed for free from many different online sources (repositories).
A repository is a place where packages are located and stored so we can install them from it. Some of the most popular repositories for R packages, are:
CRAN: Comprehensive R Archive Network(CRAN) is the official R repository.
Github: Github is the most popular repository for open source projects.
Bioconductor: Bioconductor is a topic-specific repository, intended for open source software for bioinformatics.
Add-on R packages extend the functionality of R by providing additional collections of R functions, sample data, and documentation for the included functions in a well-defined format.
To use an add-on package we need to:
Install the package from a repository. Once we’ve installed a package, we don’t need to install it again unless we want to update it.
Load the package in R session. Add-on packages are not loaded by default when we start an R session in RStudio. Each add-on package needs to be loaded explicitly every time we start RStudio.
For example, among the many add-on packages, we will use in this textbook are the dplyr package for data wrangling, the ggplot2 package for data visualization and and the rstatix package for statistical tests.
Let’s now show how to perform these two steps for the `rstatix`` package for data visualization.
5.2 Package installation
There are two ways to install an add-on R package as follows:
A. Installing packages using RStudio interface
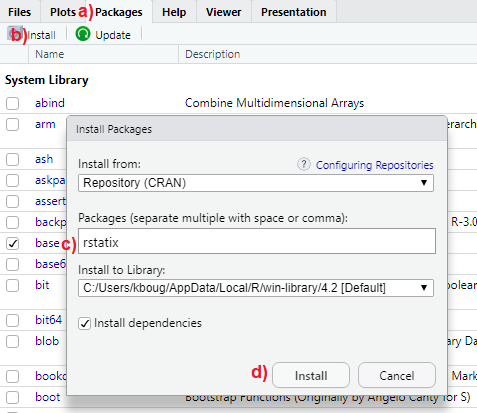
Let’s install the rstatix package as shown in Figure Figure 5.1. In the output pane of RStudio:
- Activate the “Packages” tab.
- Click on “Install”.
- Type the name of the package
rstatixin the box under “Packages (separate multiple with space or comma):”. - Press “Install.”
B. Installing packages from repositories using command
For installing the rstatix package from CRAN we type the following command in the Console pane of RStudio and press Enter on our keyboard:
Note we must include the quotation marks around the name of the package. In order to install several package at once, we just have to use the install.packages()` function as follows:
install.packages(c("rstatix", "dplyr", "ggplot2"))We only have to install a package once. However, if we want to update an installed package to a newer version, we need to re-install it using the previous command.
Moreover, suppose, for instance, that we want to download the development version of the rstatix package from GitHub. The first step is to install and load the devtools package, available on CRAN. On Windows, in case we encounter some error, we might need to install the Rtools program. Then we can call the install_github() function to install the R package from GitHub.
In case we need to install an older version of a package the simplest method is to use the provided install_version() function of the devtools package to install the version we need.
5.3 Package loading
After we’ve installed a package, we need to load it in the current R session using the library() command (note that the the quotation marks are not necessary when we are loading a package). For example, to load the rstatix package, we run the following code in the console pane:
Packages Vs Libraries: There is always confusion between a package and a library. The directories in R where the packages are stored are called the libraries.
If the cursor is blinking next to the > prompt in Console, the rstatix package was successfully installed and it is ready for use; otherwise, we get the following error:
Error in library(rstatix) : there is no package called ‘rstatix’
If we forget to load rstatix in our R session, when we try to use the functions included in this package such as t_test(), we will get an error :
Error in … : could not find function t_test
There is one way in R that can use a function without using library(). To do this, we can simply use the notation package::function .
For example:
rstatix::t_test()The above notation tells R to use the t_test function from rstatix without load the rstatix package.
5.4 The {tidyverse} package
In this textbook we will use the tidyverse package. Actually, the tidyverse is a collection of R packages designed for data science that work in harmony.
The command install.packages("tidyverse") will install the complete tidyverse. The tidyverse package provides a shortcut for downloading the following packages:
[1] "broom" "conflicted" "cli" "dbplyr"
[5] "dplyr" "dtplyr" "forcats" "ggplot2"
[9] "googledrive" "googlesheets4" "haven" "hms"
[13] "httr" "jsonlite" "lubridate" "magrittr"
[17] "modelr" "pillar" "purrr" "ragg"
[21] "readr" "readxl" "reprex" "rlang"
[25] "rstudioapi" "rvest" "stringr" "tibble"
[29] "tidyr" "xml2" "tidyverse" When we load the tidyverse package with the command library(tidyverse), R will load the nine core packages namely dplyr,forcats, ggplot2, lubridate, purrr, readr, stringr, tibble, and tidyr, and make them available in our current R session.
The packages out of the core list of tidyverse have more specialized usage and are not loaded automatically with the command library(tidyverse). So, we need to load each one explicitly with the library() function if we want to use them.
5.5 The {here} package
When we work with Projects in RStudio we may find useful the here package. The main function of the package, here(), builds a path relative to the top level of our RStudio project every time we call it. It allows us to navigate throughout each of the sub-folders and files in our project using relative paths.
We can think of the paths as directions to the files. There are two kinds of paths: absolute paths and relative paths.
For example, suppose that Emily and Paul are working on a project together and want to read in R studio of their computers an excel file named covid19.xlsx that is stored within a “data” folder. They could do this using either an absolute or a relative path as follows:
A. Reading data using an absolute path
Emily’s file is stored at C:/emily/project/data/covid19.xlsx and the command in R should be:
library(readxl)
dat <- read_excel("C:/emily/project/data/covid19.xlsx") while Paul’s file is stored at C:/paul/project/data/covid19.xlsx and the command in R should be:
library(readxl)
dat <- read_excel("C:/paul/project/data/covid19.xlsx")Even though Emily and Paul stored their files in their “C” disk, the absolute paths are different due to their different usernames.
B. Reading data using a relative path
For a relative path the command in R should be:
library(readxl)
dat <- read_excel(here("data", "covid19.xlsx"))In this case, here() tells R that the folder structure starts at the project-level. The relative path from inside the project folder (data/covid19.xlsx) is the same on both computers; any code that uses relative paths will work on both computers.